Homepage¶
Welcome to Brainnetome DiffusionKit’s Homepage¶
Note
- a full pipeline for (pre-)processing and visualization for diffusion MRI data.
- cross-platform support and a small installation size without 3rd party dependency.
- utilize a graphical interface along with command-line programs that enables easy operation and batch processing.
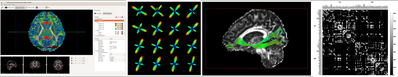
Brainnetome DiffusionKit is a light one-stop cross-platform solution to dMRI data analysis. The package delivers a complete pipeline from data format conversion to preprocessing, from local reconstruction to fiber tracking, and from fiber statistics to visualization.
It was developed as a cross-platform framework, using VTK [2] for visualization, and Qt for GUI design. Both GPU and CPU computing were implemented for visualization to achieve high frame-rate, for rendering complex scene like whole brain tractographs in particular. The project was managed using the compiler-independent CMake [3], which is compatible with gcc/g++ and MS Visual Studio, etc. Well-established algorithms, such as the DICOM conversion tool dcm2nii by Chris Rorden [4] and the constrained spherical deconvolution (CSD) for HARDI reconstruction in MRtrix [5], were adopted with improved interface and user experience.
- Visit Manual page for a complete list of usage instructions.
- Visit Tutorial page to take a look at how DiffusionKit can solve your practical problems.
- Visit Screenshot page to see how the GUI front-end looks like.
- Visit Support page to submit comments, or email us at diffusion.kit@nlpr.ia.ac.cn .
Please see the navigation sidebar to the left to begin.
Important
DiffusionKit v1.5 Released! Please Update your DiffusionKit.
- Fix bugs.
- Add features in 2D image display.
- Add functions to transform fiber track format between DiffusionKit and MRTrix.
The citation for DiffusionKit could be:
Sangma Xie, Liangfu Chen, Nianming Zuo and Tianzi Jiang, DiffusionKit: A Light One-Stop Solution for Diffusion MRI Data Analysis, Journal of Neuroscience Methods, vol. 273, pp. 107-119, 2016. [PDF] .
Note
Document last updated on Apr 27, 2020.